A research by the Computational Biology Laboratory of the CIC bioGUNE -member of the Basque Research & Technology Alliance, BRTA- and the Luxembourg Center for Systems Biomedicine (LCSB) has managed to describe a new method to identify the molecules that amplify and maintain the inflammatory response to infection. The work has been published in the jprestigious Science Advances journal. The team of researchers uses their novel method to identify these molecules in patients with Covid-19 and thus propose new potential targets for therapeutic intervention for severe cases of the disease.
When infected by a virus, the human immune system is often activated to fight the intruder. The immune response is produced from a series of complex processes that involve various types of cells that communicate with each other through proteins called cytokines. One of the main tasks of the immune system is to identify and eliminate cells in the body that have become infected and cause inflammation.
"Normally, the immune system is highly controlled and manages to reduce tissue damage as much as possible", explains Antonio del Sol, Ikerbasque researcher, and group leader at the CIC bioGUNE Computational Biology Laboratory. "However, under conditions in which the pathogen factor cannot be easily eliminated, we know that patients can develop a hyperinflammatory condition in which the immune response is amplified and maintained by positive feedback loops. This hyper-inflammatory state increases the severity of symptoms and can even lead to death. The current challenge is how to alleviate inflammation without diminishing the patient's ability to eliminate SARS-CoV-2, the virus that causes Covid-19" he adds.
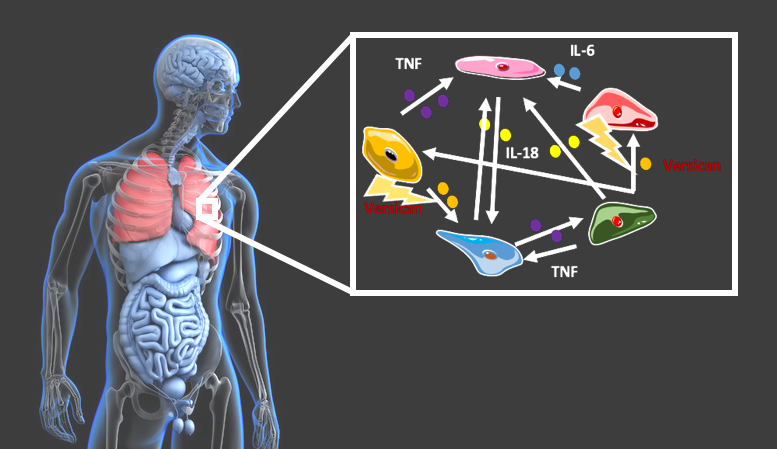
To solve this problem, researchers have developed a new method that allows the identification of modulators of the immune response. Once the molecules that amplify inflammation have been identified, their activity can be reduced to safe levels. By analyzing more than 1,700 cell-to-cell interactions, the group created a comprehensive map of the immune response in the lungs of Covid-19 patients with mild and severe symptoms. The special feature of this approach is that the genomic information of individual cells was used to create these maps, thus also representing the different cell types of the immune system.
"We decided to use our model in the context of Covid-19, as patients with severe disease courses often have hyperinflammation, which is life-threatening" says Sascha Jung, first author of the study. "We found that increased inflammation is caused by feedback mechanisms, which also partially damage immune cells and thus exacerbate the inflammatory response" he says.
Prof. Antonio Del Sol and his team used their model to identify those molecules that might be capable of modulating the immune system's runaway inflammatory response. Del Sol summarizes the results of this approach as follows: "By computationally disrupting the genes involved in feedback loops, we deduced that the Versican proteins and the Toll-2-like receptor play an important role. According to our model, the inhibition of either of these two molecules can alter up to 75% of the feedback loops without interfering with the overall immune response".
The Versican protein resides in the extracellular matrix and participates in various intercellular communication processes. For its part, the Toll-2 receptor resides on the surface of cells, where it detects pathogenic molecules. Both proteins are known to play a role in the human immune response.
The results of this study have already caught the attention of immunologists at the "Cytokines" conference, the most important in the world on basic, translational and clinical research related to the biology of cytokines, where Del Sol's work has been presented. According to these novel discoveries, the predicted proteins constitute a possible target for medical intervention in the treatment of Covid-19, an option that is currently being evaluated in animal models by immunologists from the University of Washington (USA).
The article can be found here: https://advances.sciencemag.org/content/7/6/eabe5735/tab-article-info
.png)
